ch3.intro_to_metaflow
Ch 3. Introducing Metaflow
The basics of Metaflow
Why was Metaflow created, and what problem does it solve?
- Origin: Developed at Netflix in 2017 to empower data scientists to independently build, deliver, and operate complete data science applications.
- Business need:
- Large companies like Netflix have numerous potential use cases.
- Rapid experimentation with minimal overhead is essential.
- Promising experiments should easily transition to production without requiring large teams.
How does Metaflow align with the four Vs of data science infrastructure?**
-
Volume:
- Supports more applications with fewer people.
- Reduces accidental complexity by providing a unified workflow-based approach.
-
Variety:
- Flexible across different types of data science applications.
- Opinionated about lower infrastructure layers (to ensure reliability), less opinionated about top layers (to support diverse use cases).
-
Velocity:
- Streamlines prototyping and production interaction.
- Uses idiomatic Python to enhance developer speed and comfort.
-
Validity:
- Enforces best practices to support production-grade robustness, even without DevOps expertise.
What is Metaflow’s role from an engineering perspective?
- Acts as a substrate for integration, not a replacement for existing tools.
- Recognizes that most companies already use solid tools (e.g., for storage, compute, ML libraries).
- Focuses on bridging and binding these components into cohesive data science workflows.
- Enables integration into business systems rather than isolation on a separate platform.
How is Metaflow designed for flexibility and extensibility?
-
Plugin architecture:
- Allows different backends for different stack layers.
- Supports a range of operations using cloud-native primitives.
- Compatible with major cloud providers (compute and storage abstractions).
-
Gentle adoption curve:
- Start on a local machine in “single-player mode.”
- Scale to cloud infrastructure incrementally as project needs grow.
Metaflow workflow
What are the key ideas for writing a basic Metaflow workflow?
- A workflow simplifies structuring data science applications by organizing logic into sequential steps, similar to notebook cells.
- Each workflow is defined as a directed acyclic graph (DAG) where steps (nodes) connect via directed edges.
- Metaflow provides structure, built-in execution control, and automatic experiment tracking.
What are the six rules for defining a Metaflow workflow?
-
Class definition:
- A workflow is a Python class derived from
FlowSpec. - Class names conventionally end with
Flow.
- A workflow is a Python class derived from
-
Defining steps:
- A step is a method with the
@stepdecorator. - Method body can contain arbitrary Python code.
- Optional docstrings can describe each step.
- A step is a method with the
-
Step execution:
- Metaflow runs each step as a separate task, an atomic unit of execution.
-
Starting step:
- The first step must be named
start.
- The first step must be named
-
Defining step transitions:
- Use
self.next(step_name)as the final line in a step to define the next step.
- Use
-
Ending step:
- The final step must be named
end, with noself.next()required.
- The final step must be named
-
Single flow per module:
- One flow per Python file.
- Instantiate the flow inside
if __name__ == '__main__':.
Hello world
from metaflow import FlowSpec, step
class HelloWorldFlow(FlowSpec):
@step
def start(self):
"""Starting point"""
print("This is start step")
self.next(self.hello)
@step
def hello(self):
"""Just saying hi"""
print("Hello World!")
self.next(self.end)
@step
def end(self):
"""Finish line"""
print("This is end step")
if __name__ == '__main__':
HelloWorldFlow()How to read and understand a Metaflow workflow?
- Locate the
startstep: Marks the beginning of the flow. - Trace execution using
self.next(): Identify the sequence of steps. - Read subsequent steps: Continue tracing until reaching the
endstep.
How to run and validate a Metaflow flow?
-
Save the flow in
helloworld.py. -
Validate structure without running steps:
python helloworld.py -
Disable linter (optional):
python helloworld.py --no-pylint -
Show DAG structure:
python helloworld.py show -
Run the flow:
python helloworld.py run
What information is shown in the run output header?
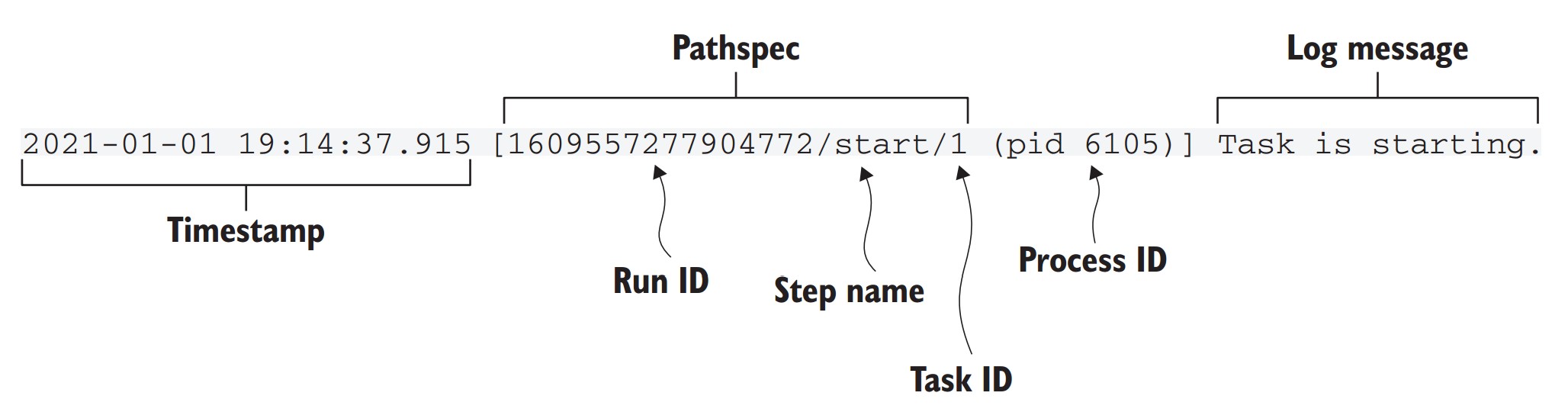
- Timestamp: When output was generated.
- Pathspec: Uniquely identifies the task (
run_id/step/task_id). - Step name: Current step being executed.
- PID (process ID): OS-level process executing the task.
- Log message: Output from Metaflow or user-defined code.
Why are unique identifiers important in Metaflow?
- Enable automatic experiment tracking.
- Help identify and compare different runs.
- Support reproducibility by tracking exact versions and results.
How to inspect logs from a specific past run?
-
Use the logs subcommand with a valid pathspec:
python helloworld.py logs <pathspec> -
Example:
python helloworld.py logs 1609557277904772/start/1 -
View available options:
python helloworld.py logs --help
### What built-in features does Metaflow provide?
- Automatic conversion of Python script into a CLI tool.
- No manual setup for command-line parsing or log management.
- Each step is a separate subprocess
- Steps resemble notebook cells but are executed as independent OS tasks.
- Experiment tracking is automatic, no additional setup needed.
- The structure ensures clarity and simplifies debugging, unlike unstructured Python scripts.
What are the key questions to determine data flow in a workflow?
-
Input access:
- How the workflow accesses raw or source data.
-
Internal state transfer:
- How intermediate (transformed) data is passed between steps.
-
Output accessibility:
- How the final results or artifacts are made available to external systems.
How does step execution determine data flow in a Metaflow workflow?
- Steps are executed in a directed order, from the
startto theend. - Data flows downstream only, from earlier steps to later ones.
- Reverse flow is not possible
Why does explicit data flow matter in workflows compared to notebooks?
-
Notebook behavior:
- Allows arbitrary execution order.
- Jupyter kernel maintains an implicit hidden state.
- Results can become unpredictable or inconsistent, depending on execution order.
-
Workflow behavior:
- Enforces explicit, ordered execution.
- Each step’s output becomes input to the next step.
- Ensures reproducibility and consistency of results.
-
Notebooks are well-suited for exploratory tasks and ad-hoc analysis.
-
For production workflows or complex pipelines, explicit data flow in a DAG structure is preferred for accuracy and maintainability.
How do workflows improve upon notebook-style development for serious applications?
- Eliminates risk of accidental re-evaluation inconsistencies.
- Forces clear dependencies between transformations.
- Ensures that all outputs are produced in a deterministic manner.
- Makes workflows easier to debug, test, and maintain.
Managing data flow
What is the challenge of managing state in distributed workflows?
- In single-process software, state can remain in memory between steps.
- In distributed workflows (e.g., using multiple computers or GPUs), in-memory state cannot be shared directly.
- Therefore, state must be transferred between steps running on different machines.
How can we transfer state between workflow steps?
- Persist the state after a step completes:
- Save relevant data (variables, intermediate results) to storage.
- Load the state before a new step starts:
- Retrieve previously saved data so the step can continue processing.
What are common implementation options for persisting state?
- Workflow frameworks often do not enforce a specific method.
- Users choose their own storage method (e.g., databases).
- Each step includes custom logic to:
- Load necessary data.
- Store output state after execution.
What are the downsides of manual persistence?
- Adds boilerplate code to every step.
- Increases cognitive load for the data scientist.
- Leads to workarounds:
- Merging unrelated operations into one step to avoid boilerplate.
- Persisting only outputs strictly needed by downstream steps.
- Workflow logic becomes harder to follow (poor readability).
- Debugging failures in production becomes more difficult.
- Missing historical context reduces observability and traceability.
What is the recommended approach?
- Use a transparent mechanism for state transfer:
- Abstracts away physical separation of computing resources.
- Minimizes boilerplate code.
- Encourage liberal persistence of data, even if not strictly required.
- Enhanced monitoring, debugging, reproducibility, and sharing of workflows.
- Storage cost is minimal relative to the cost of a data scientist’s time.
- Avoid redundant input data storage—addressed in detail in Chapter 7.
- Workflow design should prioritize logical clarity over implementation shortcuts.
Metaflow Artifacts
How does Metaflow make data flow nearly transparent to the user?
- Automatic persistence of all instance variables (
self.var) in each step. - These persisted instance variables are called artifacts.
- Artifacts can be any serializable Python object (via
pickle). - Stored in a central datastore managed by Metaflow.
What is the lifecycle of an artifact in Metaflow?
- Step completion:
- All instance variables are serialized and stored as immutable artifacts in the datastore.
- Each artifact is permanently associated with its producing step via a pathspec.
- Subsequent steps:
- May read artifacts from previous steps.
- May create new versions, but never overwrite previous ones.
Example: CounterFlow using persisted state
from metaflow import FlowSpec, step
class CounterFlow(FlowSpec):
@step
def start(self):
self.count = 0
self.next(self.add)
@step
def add(self):
print("The count is", self.count, "before incrementing")
self.count += 1
self.next(self.end)
@step
def end(self):
self.count += 1
print("The final count is", self.count)
if __name__ == '__main__':
CounterFlow()self.count = 0is initialized instart.- Incremented in
add→ persisted as a new artifact. - Incremented again in
end. - Run using:
python counter.py run
What happens under the hood when managing state?
- Each task runs in its own process, possibly on a different computer.
- Snapshotting occurs at the end of each step:
- All
selfvariables are serialized and stored. - Enables resuming, debugging, and cross-machine execution.
- All
How can you inspect artifacts?
- Use the
dumpcommand to inspect step outputs:python counter.py dump <run_id>/end/3 - Use the Metaflow Client API for programmatic access (covered later in section 3.3.2).
Rule of thumb for using instance variables
- Use
self.variablefor:- Any data that must persist across steps.
- Any data that may be helpful for debugging.
- Use local variables for:
- Temporary, within-step computations.
How does Metaflow’s datastore function internally?
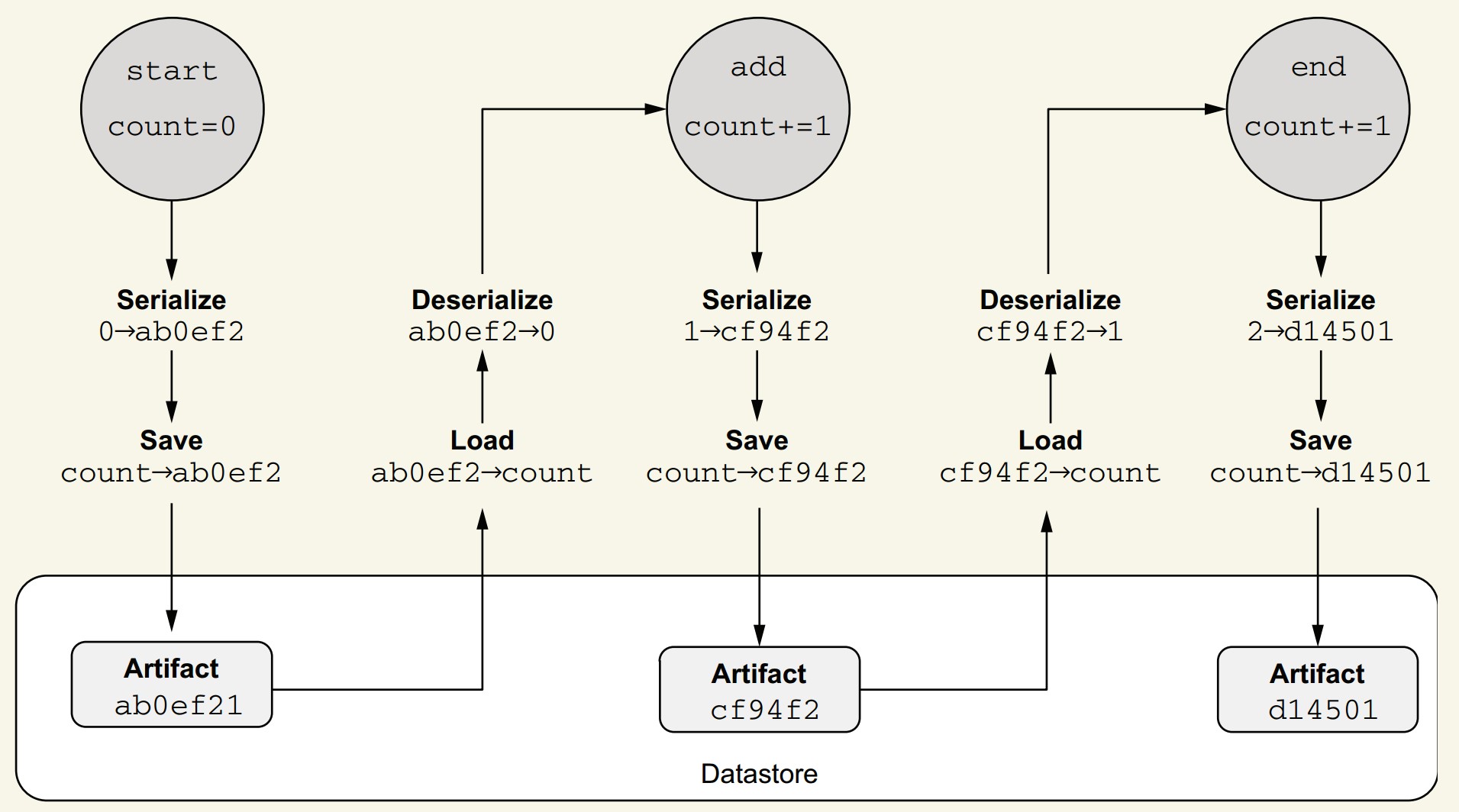
- After task execution:
- Metaflow inspects instance variables.
- Each is serialized (currently with
pickle) into a byte blob (e.g.,ab0ef2).
- Stored as immutable artifacts:
- Linked to step metadata (e.g.,
start -> ab0ef2).
- Linked to step metadata (e.g.,
- When accessed by another step:
- Artifact is fetched from the datastore using metadata and flow order.
- Datastore is content-addressed (similar to Git):
- Artifacts are hashed by content.
- Automatic deduplication prevents redundant storage.
Benefits of this approach
- Minimizes user effort in managing state explicitly.
- Enables reproducibility, observability, and debugging.
- Efficient storage without user intervention.
- Artifact versioning enables easy tracking and auditing of workflow runs.
More on step
What makes a good step in a Metaflow workflow?
- Think of steps as checkpoints:
- Artifacts are persisted at the end of each step.
- Workflow execution can be resumed from any completed step.
- Artifacts are available for inspection after step completion.
Why keep steps reasonably small?
- Improves resilience:
- Less work lost in case of failure.
- Enhances observability:
- You can monitor progress and intermediate state more frequently.
- Enables fine-grained debugging:
- Narrow down where issues occur.
- Supports incremental development:
- Easier to build and test workflows in parts.
Why not make steps too small?
- Each step incurs overhead:
- Task launching.
- Artifact persistence.
- Too many tiny steps may:
- Slow down the workflow.
- Create unnecessary complexity.
- If overhead becomes apparent, you can merge small steps later.
How does step size affect code readability?
- Use
python flow.py showto inspect the workflow graph. - Large steps:
- May hide logical structure.
- Can make the graph less meaningful.
- Smaller, focused steps:
- Improve code clarity.
- Make workflow structure more comprehensible.
Rule of thumb
- Organize workflows into logical, explainable steps.
- When uncertain:
- Favor smaller steps—easier to understand and debug.
- You can always refactor and combine them if needed.
Metaflow parameters
How do you pass data into a Metaflow flow?
- Use Parameters to pass external values at the time of running the flow.
- Unlike artifacts, which pass data between steps, Parameters provide data to the flow at the start.
What are Metaflow Parameters?
- Special class-level constructs that:
- Are immutable once the run starts.
- Are available to all steps, including
start. - Are treated as artifacts, so they're tracked and inspectable.
- Parameters simplify experiment tracking by making runs reproducible and traceable.
- All parameters are recorded like other artifacts, aiding future inspection and debugging.
How do you define a Parameter?
Use the Parameter() constructor at the class level. Four main elements are required:
- Instantiate
Parameterat class scope. - Assign it to a variable (e.g.,
animal,count). - Specify a name shown to users (e.g.,
'creature','count'). - Set a type:
- Either via
type=...(e.g.,float,int) - Or by providing a default value (type is inferred)
- Either via
Example:
from metaflow import FlowSpec, Parameter, step
class ParameterFlow(FlowSpec):
animal = Parameter('creature', help="Specify an animal", required=True)
count = Parameter('count', help="Number of animals", default=1)
ratio = Parameter('ratio', help="Ratio between 0.0 and 1.0", type=float)
@step
def start(self):
print(self.animal, "is a string of", len(self.animal), "characters")
print("Count is an integer: %s+1=%s" % (self.count, self.count + 1))
print("Ratio is a", type(self.ratio), "whose value is", self.ratio)
self.next(self.end)
@step
def end(self):
print("done!")
if __name__ == '__main__':
ParameterFlow()How do you run a flow with parameters?
- Required parameters must be specified via CLI:
python parameters.py run --creature seal- Optional parameters use defaults unless specified:
python parameters.py run --creature seal --count 10 --ratio 0.3- Metaflow automatically converts values to specified types.
Can you change a parameter’s value in your code?
- No — parameters are immutable.
- To manipulate a parameter, copy it to another artifact:
self.mutable_count = self.count
self.mutable_count += 1How to avoid specifying parameters repeatedly?
- Set parameters using environment variables:
export METAFLOW_RUN_CREATURE=dinosaur
python parameters.py run --ratio 0.25- CLI values override environment variables:
python parameters.py run --creature otter --count 10 --ratio 0.3creatureis now set to"otter"instead of"dinosaur".
Complex parameters
How do you define complex parameters in Metaflow?
- Basic parameters support scalar types:
str,int,float,bool. - For more complex structures (e.g., lists, dicts), use JSON-encoded parameters.
How do JSON-encoded parameters work?
- JSON values are passed as strings on the command line.
- Use the
JSONTypetype from Metaflow for decoding.
from metaflow import FlowSpec, Parameter, step, JSONType
class JSONParameterFlow(FlowSpec):
mapping = Parameter('mapping',
help="Specify a mapping",
default='{"some": "default"}',
type=JSONType)
@step
def start(self):
for key, value in self.mapping.items():
print('key', key, 'value', value)
self.next(self.end)
@step
def end(self):
print('done!')-
Run with:
python json_parameter.py run --mapping '{"mykey": "myvalue"}' -
Use single quotes to prevent shell issues with special characters.
How do you handle large JSON parameters?
-
Store JSON in a file (e.g.,
myconfig.json):echo '{"largekey": "largevalue"}' > myconfig.json -
Pass using shell substitution:
python json_parameter.py run --mapping "$(cat myconfig.json)"
How do you include actual files as parameters?
- Use
IncludeFilefor small/medium-size files (e.g., CSV, JSON). - Artifacts created via
IncludeFileare snapshotted and versioned.
from metaflow import FlowSpec, Parameter, step, IncludeFile
from io import StringIO
import csv
class CSVFileFlow(FlowSpec):
data = IncludeFile('csv',
help="CSV file to be parsed",
is_text=True)
delimiter = Parameter('delimiter',
help="delimiter",
default=',')
@step
def start(self):
fileobj = StringIO(self.data)
for i, row in enumerate(csv.reader(fileobj, delimiter=self.delimiter)):
print("row %d: %s" % (i, row))
self.next(self.end)
@step
def end(self):
print('done!')-
Create a test CSV:
echo "first,second,third\na,b,c" > test.csv -
Run with:
python csv_file.py run --csv test.csv
Why use IncludeFile instead of reading files directly?
- Files passed via
IncludeFilebecome immutable artifacts:- Versioned alongside the flow run.
- Available even if the original file is deleted or changed later.
- Supports reproducibility and traceability of input data.
Miscellaneous Notes
- Use
JSONTypefor structured config-like inputs. - Use
IncludeFilefor datasets or larger structured inputs. IncludeFileis not meant for very large files—chapter 7 covers scalable data handling.
Branches
What is the role of branches in Metaflow workflows?
- Branches allow concurrent execution of steps.
- Enables efficient use of resources like multicore CPUs or distributed clusters.
- Provides a simple abstraction of parallelism, easier than low-level concurrency models like multithreading.
When should you use branches?
- Use branches when steps can run independently based on data flow:
- If step A produces data needed by both B and C, but B and C don’t need each other’s output → branch B and C from A.
- Join their outputs later in a common step D if needed.
- Example DAG:
- A (fetch data)
↓ - B (train model 1) & C (train model 2) — run in parallel
↓ - D (compare and choose best model)
- A (fetch data)
What are the benefits of branching?
- Performance: Steps B and C execute in parallel.
- Clarity: Reflects the true data flow and step dependencies.
- Maintainability: Easier to understand and reason about.
Rule of thumb
When two or more steps can be executed independently, define them as parallel branches.
Why not automate parallelization?
- Automatic parallelism is hard with arbitrary Python:
- Python code often lacks enough structure for safe auto-parallelization.
- Steps may use external libraries or services.
- Manual control leads to better clarity and fewer errors.
- Workflows are also communication tools, meant to be human-readable.
What are valid DAG structures in Metaflow?
- Split step: A step that initiates branching (e.g., A).
- Join step: A step that merges branches back together (e.g., D).
- Every
splitmust have a correspondingjoin.
Rules for DAG validity:
- Every
splitmust be matched by ajoin(like matching parentheses). - A
joinstep can only join steps that share a commonsplitparent.
- Invalid: Joining steps that came from different splits.
- Valid: All joined branches must originate from the same split step.
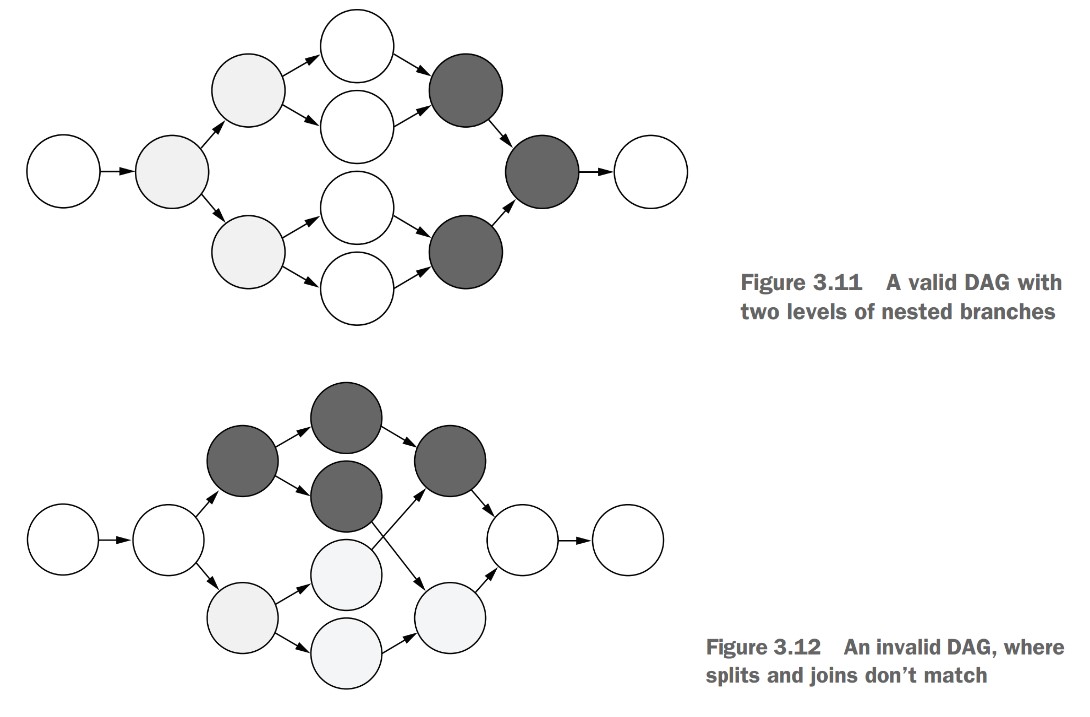
Why are these constraints necessary?
- Metaflow’s DAG constraints ensure clean, reproducible parallel workflows.
- Branching is a core feature to speed up processing and clarify structure.
- The constraints prevents ambiguous artifact lineage.
- Keeps DAGs clear and readable:
- In a valid DAG, no arrows (edges) have to cross.
- Simplifies tracking and debugging data flow across steps.
Static branch
What is a static DAG in Metaflow?
- A static DAG has a fully defined structure before execution begins.
- All branches and joins are declared explicitly using
self.next(...).
How is branching defined in a static DAG?
- Split step: Use
self.next(step1, step2, ...)to define parallel branches. - Join step: Add an extra
inputsparameter to access upstream branches.
Why do we need to merge artifacts after a branch?
- Branches create divergent data flows—artifacts may have different values in each branch.
- Join steps act as barriers: downstream steps can't access any upstream artifacts unless explicitly reassigned or merged.
- Even unchanged artifacts (like
creature) must be explicitly reassigned in the join step to make them available downstream.
Example: CounterBranchFlow with static branches and manual merging
from metaflow import FlowSpec, step
class CounterBranchFlow(FlowSpec):
@step
def start(self):
self.creature = "dog"
self.count = 0
self.next(self.add_one, self.add_two)
@step
def add_one(self):
self.count += 1
self.next(self.join)
@step
def add_two(self):
self.count += 2
self.next(self.join)
@step
def join(self, inputs):
self.count = max(inp.count for inp in inputs)
print("count from add_one", inputs.add_one.count)
print("count from add_two", inputs.add_two.count)
self.creature = inputs[0].creature # manual reassignment
self.next(self.end)
@step
def end(self):
print("The creature is", self.creature)
print("The final count is", self.count)
if __name__ == '__main__':
CounterBranchFlow()-
Run with:
python counter_branch.py run -
Comment out the
self.creaturereassignment injointo see the flow fail due to a missing artifact. -
You can access branches in the join step via:
inputs[0](first branch)inputs.branch_namefor inp in inputs(for aggregation withsum,max, etc.)
-
Metaflow executes branches in parallel processes, enabling true concurrency on multicore systems.
How does merge_artifacts() help?
- Automatically reassigns non-divergent artifacts.
- Replaces boilerplate like
self.creature = inputs[0].creature. - Must be called after manually merging divergent artifacts like
count.
self.merge_artifacts(inputs)How to exclude specific artifacts from merging?
- Use the
excludeoption for artifacts you don’t want to propagate downstream:
self.merge_artifacts(inputs, exclude=['increment'])- Useful for internal or debug-only variables.
Example:
from metaflow import FlowSpec, step
class CounterBranchHelperFlow(FlowSpec):
@step
def start(self):
self.creature = "dog"
self.count = 0
self.next(self.add_one, self.add_two)
@step
def add_one(self):
self.increment = 1
self.count += self.increment
self.next(self.join)
@step
def add_two(self):
self.increment = 2
self.count += self.increment
self.next(self.join)
@step
def join(self, inputs):
self.count = max(inp.count for inp in inputs)
print("count from add_one", inputs.add_one.count)
print("count from add_two", inputs.add_two.count)
self.merge_artifacts(inputs, exclude=['increment']) # !!!!!!!!!!!
self.next(self.end)
@step
def end(self):
print("The creature is", self.creature)
print("The final count", self.count)
if __name__ == '__main__':
CounterBranchHelperFlow()- Run with:
python counter_branch_helper.py run
What happens if you don’t merge artifacts properly?
- Metaflow raises an error if:
- A divergent artifact is left unmerged.
- A downstream step accesses an artifact not reassigned in the join step.
Summary of branching and merging in Metaflow:
- Use
self.next(...)for static branching. - Join step must handle all artifacts—either by:
- Manual reassignment
merge_artifacts(), with or withoutexclude
- Branching enables parallelism, but merging ensures correct and consistent data flow.
Dynamic branch
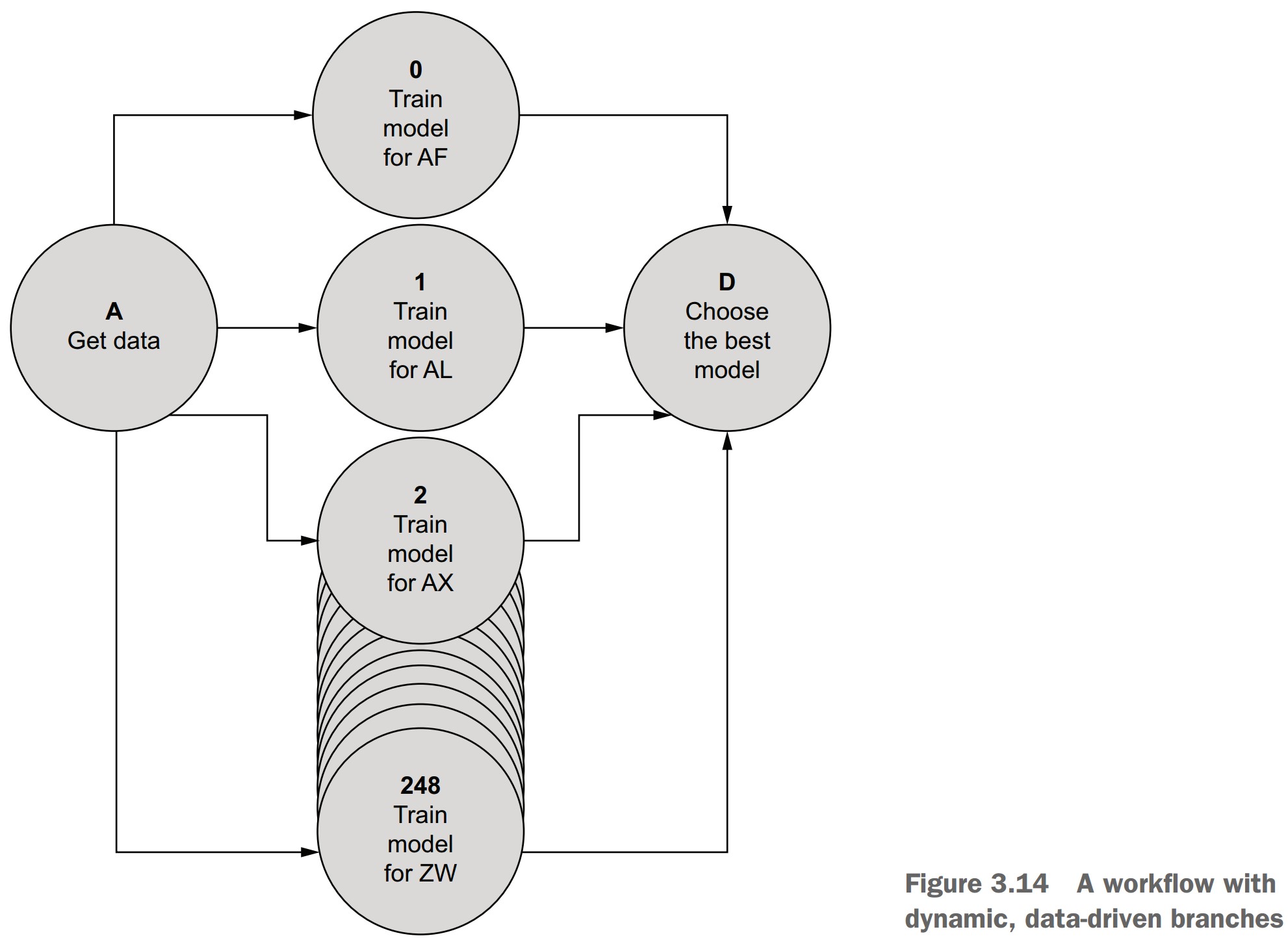
What is dynamic branching in Metaflow?
- Dynamic branches are created at runtime based on data values.
- Implemented using the
foreachconstruct. - Commonly used for data parallelism:
- Running the same step multiple times with different data (e.g., training models per country).
- Unlike static branches, where each branch runs different code.
When should you use dynamic branches?
- When the same operation needs to be applied to each item in a list.
- Typical use cases:
- Parallel training, scoring, evaluation
- Hyperparameter tuning
- Batch data processing
Syntax for a foreach split
- Use
self.next(step_name, foreach='list_artifact') - The referenced artifact must be a Python list
from metaflow import FlowSpec, step
class ForeachFlow(FlowSpec):
@step
def start(self):
self.creatures = ['bird', 'mouse', 'dog']
self.next(self.analyze_creatures, foreach='creatures')
@step
def analyze_creatures(self):
print("Analyzing", self.input)
self.creature = self.input
self.score = len(self.creature)
self.next(self.join)
@step
def join(self, inputs):
self.best = max(inputs, key=lambda x: x.score).creature
self.next(self.end)
@step
def end(self):
print(self.best, 'won!')
if __name__ == '__main__':
ForeachFlow()- Run the flow:
python foreach.py run
Key concepts for foreach steps
self.inputis available inside theforeachstep:- It contains one item from the input list.
- Must be copied to another artifact if you need to access it later.
- All iterations of the
foreachstep run in parallel.
Merging after foreach
joinstep works the same as in static branching:- Receives
inputs, a list of branches. - You can use aggregation patterns, e.g.,
max(),sum(), etc.
- Receives
- Use
merge_artifacts(inputs)to merge non-divergent values.
Parallelism patterns and foreach
- The
foreachpattern maps to common parallel computing models:- MapReduce
- Bulk Synchronous Parallel
- Fork-Join
- Parallel Map
- It matches how tools like TensorFlow or PyTorch optimize computation at a lower level.
Philosophy of Metaflow
- Metaflow focuses on human-centric application structure.
- Delegates machine-level optimization (like matrix parallelism) to ML libraries.
- Encourages defining clear workflows with data parallelism handled through
foreach.
Miscellaneous Notes
- You can see unique task identifiers in logs like
analyze_creatures/2. - These help track individual branches and their outputs.
- Dynamic branching is ideal for scalable, repeatable operations on datasets.
Control concurrency
How does Metaflow control concurrency in foreach branches?
foreachcan create thousands of tasks from a list dynamically.- By default, Metaflow limits the number of parallel splits and concurrent tasks to protect your system.
What are the two key concurrency control options?
1. --max-num-splits
- Controls the maximum number of tasks a
foreachcan spawn. - Prevents accidentally spawning massive numbers of tasks.
- Default limit: 100
- Override with:
python wide_foreach.py run --max-num-splits 10000 - Or set an environment variable:
export METAFLOW_RUN_MAX_NUM_SPLITS=100000
2. --max-workers
-
Controls how many tasks run concurrently (processes).
-
Default limit: 16
-
Can be set lower (e.g., to save memory) or higher (if your system allows):
python wide_foreach.py run --max-num-splits 1000 --max-workers 8
Example: WideForeachFlow with 1,000 tasks
from metaflow import FlowSpec, step
class WideForeachFlow(FlowSpec):
@step
def start(self):
self.ints = list(range(1000))
self.next(self.multiply, foreach='ints')
@step
def multiply(self):
self.result = self.input * 1000
self.next(self.join)
@step
def join(self, inputs):
self.total = sum(inp.result for inp in inputs)
self.next(self.end)
@step
def end(self):
print('Total sum is', self.total)
if __name__ == '__main__':
WideForeachFlow()- Run (will fail without override):
python wide_foreach.py run - Fix with:
python wide_foreach.py run --max-num-splits 1000
How does Metaflow execute these tasks?
- Uses a local scheduler:
- Converts workflow steps to tasks (processes).
- Manages execution order and concurrency.
- Respects
--max-workersas the concurrency cap.
- Tasks are queued if they exceed
--max-workers.
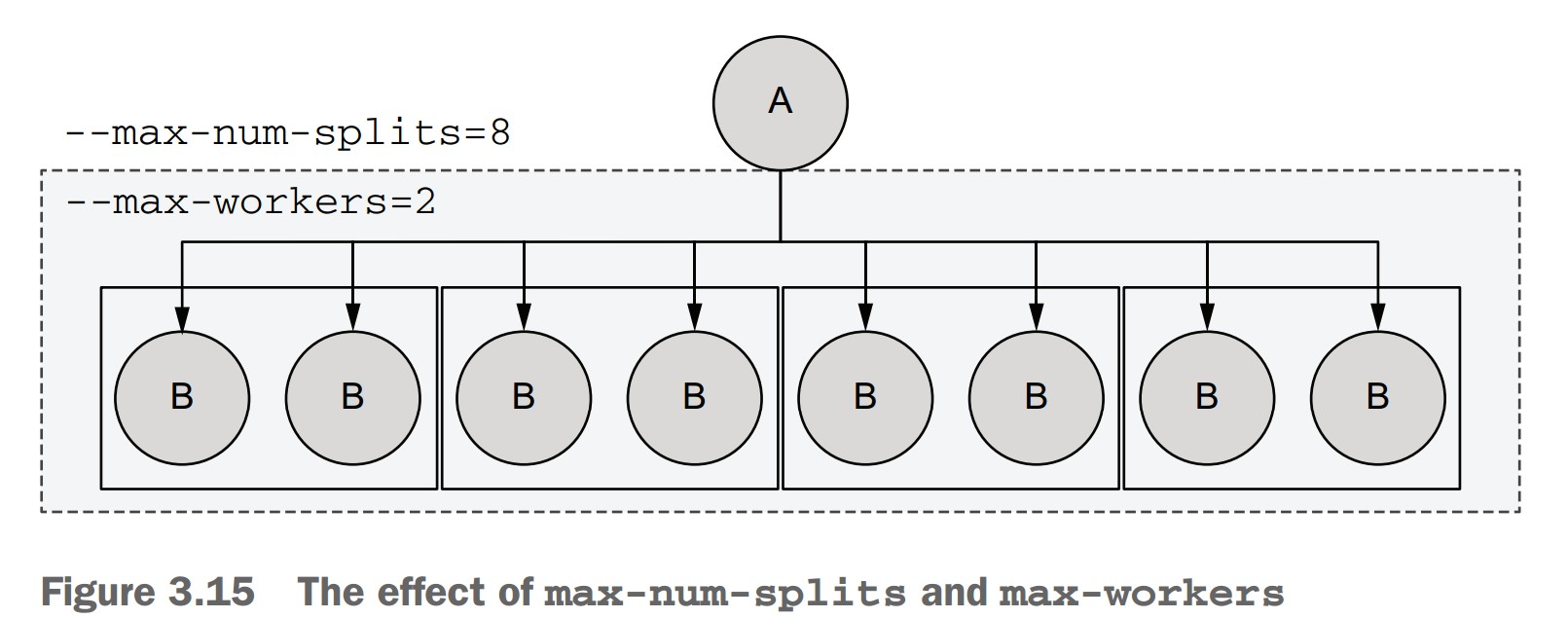
-
If a
foreachsplits into 8 tasks:--max-num-splits=8allows it.--max-workers=2means only 2 tasks run at once.- Tasks will run in 4 mini-batches of 2.
-
These limits do not affect workflow logic, only execution.
-
Helps safely test large-scale workflows locally.
-
Metaflow’s scheduler handles resource management automatically during runs.
Practical tips
- Set
--max-workersto match or stay below your CPU core count. - Lower it further if tasks use significant memory or GPU resources.
- Use
timeto benchmark performance under different settings:time python wide_foreach.py run --max-num-splits 1000 --max-workers 8 time python wide_foreach.py run --max-num-splits 1000 --max-workers 64
Starting a new project
How can you approach a new data science project using workflows?
- Use the spiral development recipe—build workflows iteratively, from outer structure to core logic.
- Helps break down the blank-file paralysis and ensures focus on key elements.
Spiral development path
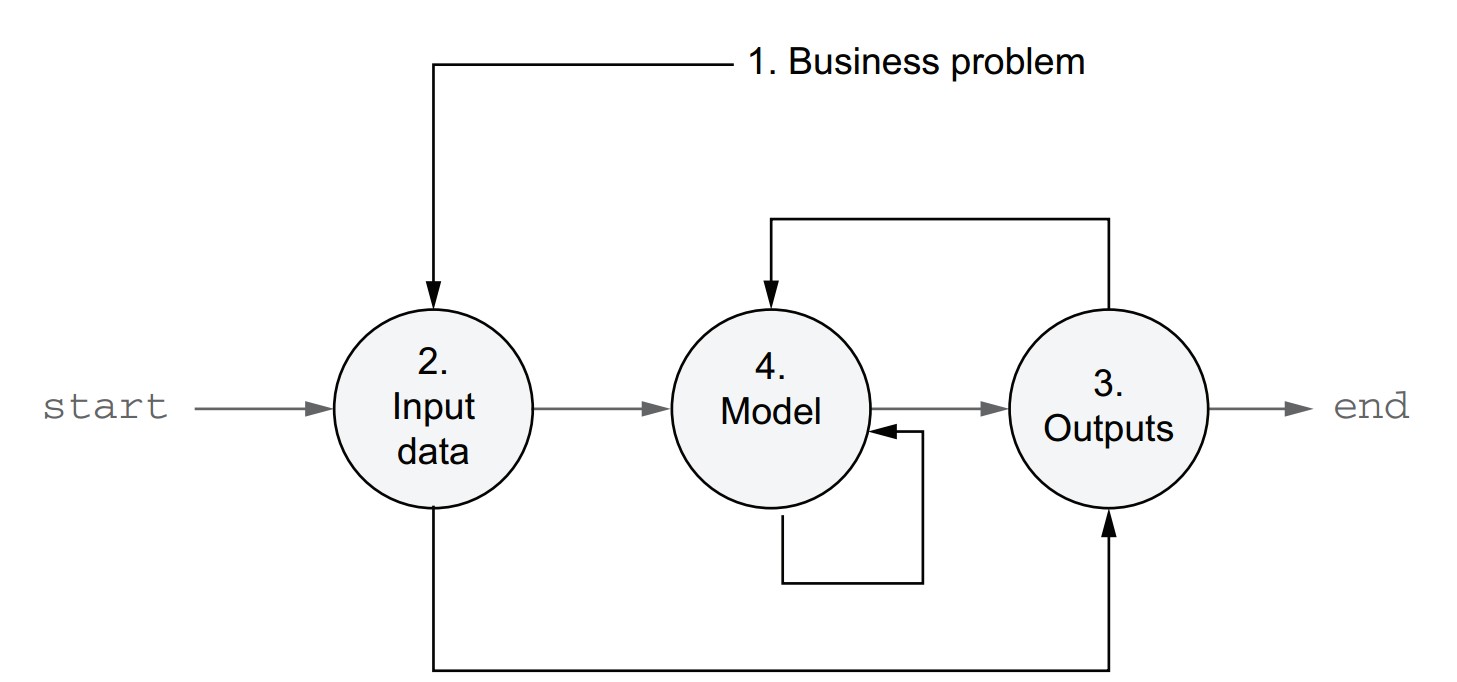
- Define the business problem you are solving.
- Identify the input data and how to read it.
- Define the desired outputs and how to store or deliver them.
- Choose initial modeling techniques to turn input into output.
- This approach ensures the problem, data, and business context guide the solution.
Why this method works well
- Clarifies objectives early to avoid building the wrong thing.
- Data prep is often underestimated—starting early saves time.
- Output integration can be complex—discovering requirements early helps.
- Start simple—a working baseline allows measurement and iterative improvement.
Infrastructure requirement
- Should support iterative development and easy experimentation.
- Metaflow is built to enable this exact pattern.
Example: Starting a new classification project
- Business problem: Classify wine types using Scikit-Learn’s wine dataset.
- Start with a minimal flow that loads and splits the data.
Code: classifier_train_v1.py
- This skeleton serves as a reusable template for future classification tasks with different datasets.
- Focus on getting a minimal version running before expanding functionality.
from metaflow import FlowSpec, step
class ClassifierTrainFlow(FlowSpec):
@step
def start(self):
from sklearn import datasets
from sklearn.model_selection import train_test_split
X, y = datasets.load_wine(return_X_y=True)
self.train_data, \
self.test_data, \
self.train_labels, \
self.test_labels = train_test_split(X, y, test_size=0.2, random_state=0)
print("Data loaded successfully")
self.next(self.end)
@step
def end(self):
self.model = 'nothingburger'
print('done')
if __name__ == '__main__':
ClassifierTrainFlow()-
Save to:
classifier_train_v1.py -
Install Scikit-Learn (if not installed):
pip install sklearn -
Run the flow:
python classifier_train_v1.py run
How to inspect artifacts to confirm data was loaded
- Use
dumpwith a real pathspec from your run:python classifier_train_v1.py dump <your_run_id>/start/1 - Output confirms artifacts exist (though large arrays may not display inline).
Notes
- In Metaflow, it's good practice to place import statements inside the steps where they are used.
- Reduces unnecessary imports and improves modularity.
- The current flow loads and splits the data; further steps (e.g., model training, evaluation) will be added incrementally.
Accessing results with the Client API
How do you access results programmatically in Metaflow?
- Use the Client API to explore and access artifacts from past runs.
- Artifacts are read-only and tied to steps, tasks, and runs.
- Helpful for:
- Debugging
- Reusing outputs
- Chaining workflows
Client API container hierarchy
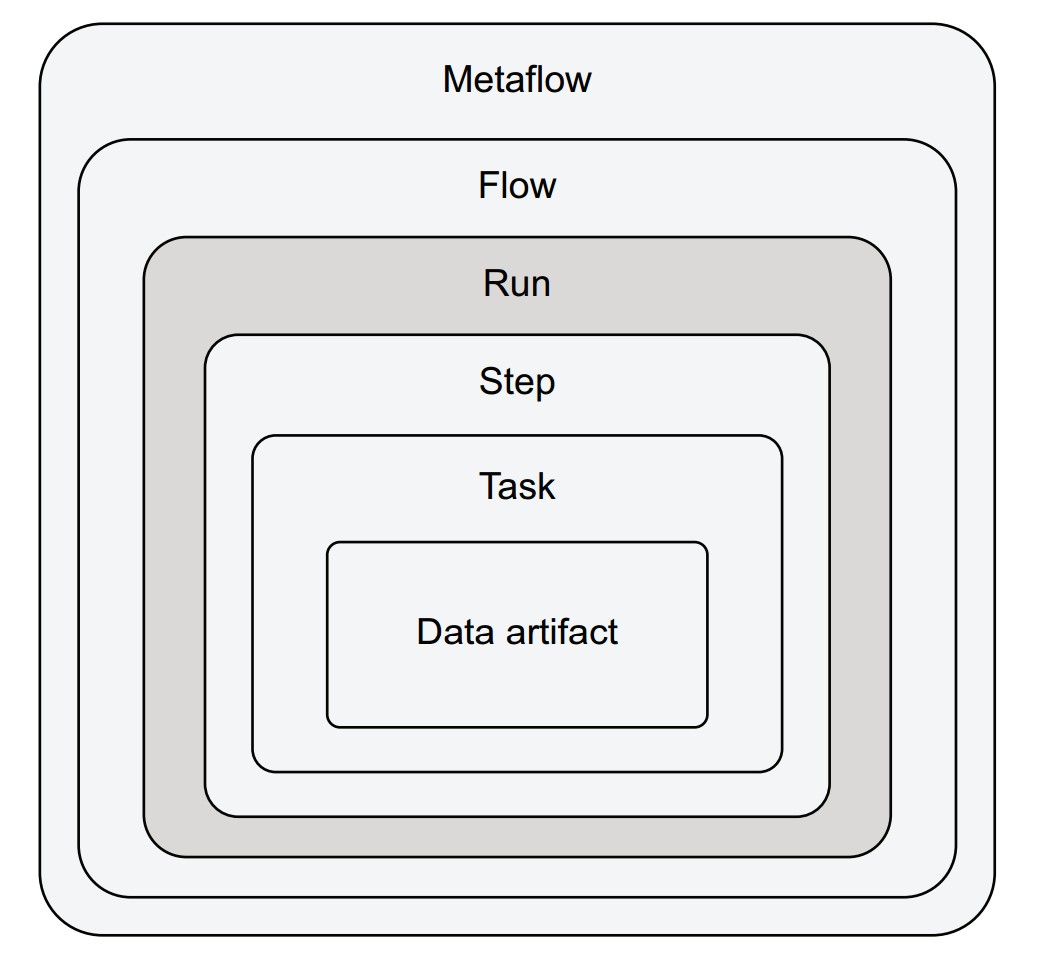
Each layer nests within the previous:
- Metaflow: All flows in the environment.
- Flow: All runs of a specific flow.
- Run: All steps executed during a single run.
- Step: All tasks in a step (one or many for
foreach). - Task: Holds artifacts and logs.
- Data artifact: A specific variable stored in a task.
Accessing artifacts via pathspec or shortcuts
-
Three ways to navigate:
from metaflow import Run # Direct pathspec run = Run("ClassifierTrainFlow/1611541088765447") # Access step and artifact run['start'].task.data.train_datafrom metaflow import Flow # Shortcut to latest run run = Flow("ClassifierTrainFlow").latest_run train_data = run['start'].task.data.train_data# Iterate over all steps of a run for step in run: print(step.id)
Recommended: Explore artifacts in a notebook
-
Open Jupyter in your flow directory:
jupyter-notebook -
In a notebook cell:
from metaflow import Flow run = Flow("ClassifierTrainFlow").latest_run run['start'].task.data.train_data # inspect artifact -
Try inspecting:
train_labelstest_datamodel
Exercise ideas
- Run the training flow again →
.latest_runwill change. - Use older runs by listing
Flow("ClassifierTrainFlow").runs. - Use
help()to inspect object properties:help(run)
Reusing artifacts in another flow
- Use the Client API to access outputs from a different flow.
- Example: Create a prediction flow that reuses the trained model from
ClassifierTrainFlow.
from metaflow import FlowSpec, step, Flow, Parameter, JSONType
class ClassifierPredictFlow(FlowSpec):
vector = Parameter('vector', type=JSONType, required=True)
@step
def start(self):
run = Flow('ClassifierTrainFlow').latest_run
self.train_run_id = run.pathspec
self.model = run['end'].task.data.model
print("Input vector", self.vector)
self.next(self.end)
@step
def end(self):
print("Model", self.model)
if __name__ == '__main__':
ClassifierPredictFlow()-
Save to:
classifier_predict_v1.py -
Run with a test vector:
python classifier_predict_v1.py run --vector '[1,2]' -
Output should print:
Model nothingburger
Notes
self.train_run_id = run.pathspeccaptures the training run’s ID.- Useful for:
- Traceability
- Debugging surprising predictions
- Auditing production results
- The Client API works across notebooks, scripts, and interactive shells.
- It’s ideal for chaining flows and building multi-stage pipelines without coupling training and inference.
Debugging failures
Common situation: failure during prototyping
-
In
ClassifierTrainFlow, we train two models in parallel: KNN and SVM. -
train_svmfails due to an invalid kernel name:svm.SVC(kernel='polynomial') # ❌ should be 'poly' -
Expected error:
ValueError: 'polynomial' is not in list
Step-by-step debugging guide
1. Find the error message
-
When running the flow:
- Error trace is printed with the failing step name (e.g.,
train_svm). - If many steps/logs are printed, use the
logscommand:python classifier_train.py logs <run_id>/train_svm/1
- Error trace is printed with the failing step name (e.g.,
-
Or use the Client API to find and inspect failed tasks:
from metaflow import Flow for step in Flow("ClassifierTrainFlow").latest_run: for task in step: if not task.successful: print("Task %s failed:" % task.pathspec) print("-- Stdout --") print(task.stdout) print("-- Stderr --") print(task.stderr) -
Optional: add filters like
if 'svm' in task.stderr:for targeted debugging.
2. Understand why the code failed
-
Investigate the error using:
-
Documentation
-
Google
-
Notebook inspection of artifacts leading up to the failure:
from metaflow import Flow run = Flow("ClassifierTrainFlow").latest_run run['train_svm'].task.data.train_data # or any other relevant artifact
-
-
Optional: Use IDE debuggers (e.g., in VS Code or PyCharm) with extra config for subprocesses.
3. Test and resume execution
-
Fix the line in
train_svm:self.model = svm.SVC(kernel='poly') # ✅ correct argument -
Instead of restarting the full run, use the
resumecommand:python classifier_train.py resume -
This resumes from the failed step and reuses successful results (e.g.,
startandtrain_knn).
Additional resume options
-
Resume from a specific step (and all following steps):
python classifier_train.py resume train_knn -
Resume from a specific past run:
python classifier_train.py resume --origin-run-id 1611609148294496 train_knn -
Parameters cannot be changed during resume (to ensure consistency with reused artifacts).
Note:
- Resumed runs receive their own unique Run ID.
- They are fully tracked and accessible via the Client API.
- Artifacts from resumed runs are consistent with their origin unless rerun steps are altered.
Final flow after the fix
from metaflow import FlowSpec, step
class ClassifierTrainFlow(FlowSpec):
@step
def start(self):
from sklearn import datasets
from sklearn.model_selection import train_test_split
X, y = datasets.load_wine(return_X_y=True)
self.train_data, self.test_data, self.train_labels, self.test_labels = \
train_test_split(X, y, test_size=0.2, random_state=0)
self.next(self.train_knn, self.train_svm)
@step
def train_knn(self):
from sklearn.neighbors import KNeighborsClassifier
self.model = KNeighborsClassifier()
self.model.fit(self.train_data, self.train_labels)
self.next(self.choose_model)
@step
def train_svm(self):
from sklearn import svm
self.model = svm.SVC(kernel='poly') # ✅ fixed
self.model.fit(self.train_data, self.train_labels)
self.next(self.choose_model)
@step
def choose_model(self, inputs):
def score(inp):
return inp.model, inp.model.score(inp.test_data, inp.test_labels)
self.results = sorted(map(score, inputs), key=lambda x: -x[1])
self.model = self.results[0][0]
self.next(self.end)
@step
def end(self):
print("Scores:")
print("\n".join("%s %f" % res for res in self.results))
if __name__ == '__main__':
ClassifierTrainFlow()What are the final steps to complete your Metaflow-based ML application?
With the training flow (ClassifierTrainFlow) now producing a valid model, the prediction flow (ClassifierPredictFlow) can be finalized with just one additional line.
Update to ClassifierPredictFlow
Add this line to the end step to make a prediction:
print("Predicted class", self.model.predict([self.vector])[0])Final version of the flow:
from metaflow import FlowSpec, step, Flow, Parameter, JSONType
class ClassifierPredictFlow(FlowSpec):
vector = Parameter('vector', type=JSONType, required=True)
@step
def start(self):
run = Flow('ClassifierTrainFlow').latest_run
self.train_run_id = run.pathspec
self.model = run['end'].task.data.model
print("Input vector", self.vector)
self.next(self.end)
@step
def end(self):
print("Model", self.model)
print("Predicted class", self.model.predict([self.vector])[0])
if __name__ == '__main__':
ClassifierPredictFlow()Test the prediction
Run the flow with a valid vector (13 attributes from the wine dataset):
python classifier_predict.py run --vector \
'[14.3,1.92,2.72,20.0,120.0,2.8,3.14,0.33,1.97,6.2,1.07,2.65,1280.0]'- Expected output:
Predicted class 0
What if the predictions seem incorrect?
- Use the Client API and Scikit-Learn’s model diagnostics:
- Inspect
.coef_,.feature_importances_, or usecross_val_score(). - Load model artifacts in a notebook for exploration.
- Inspect
Summary of the final architecture
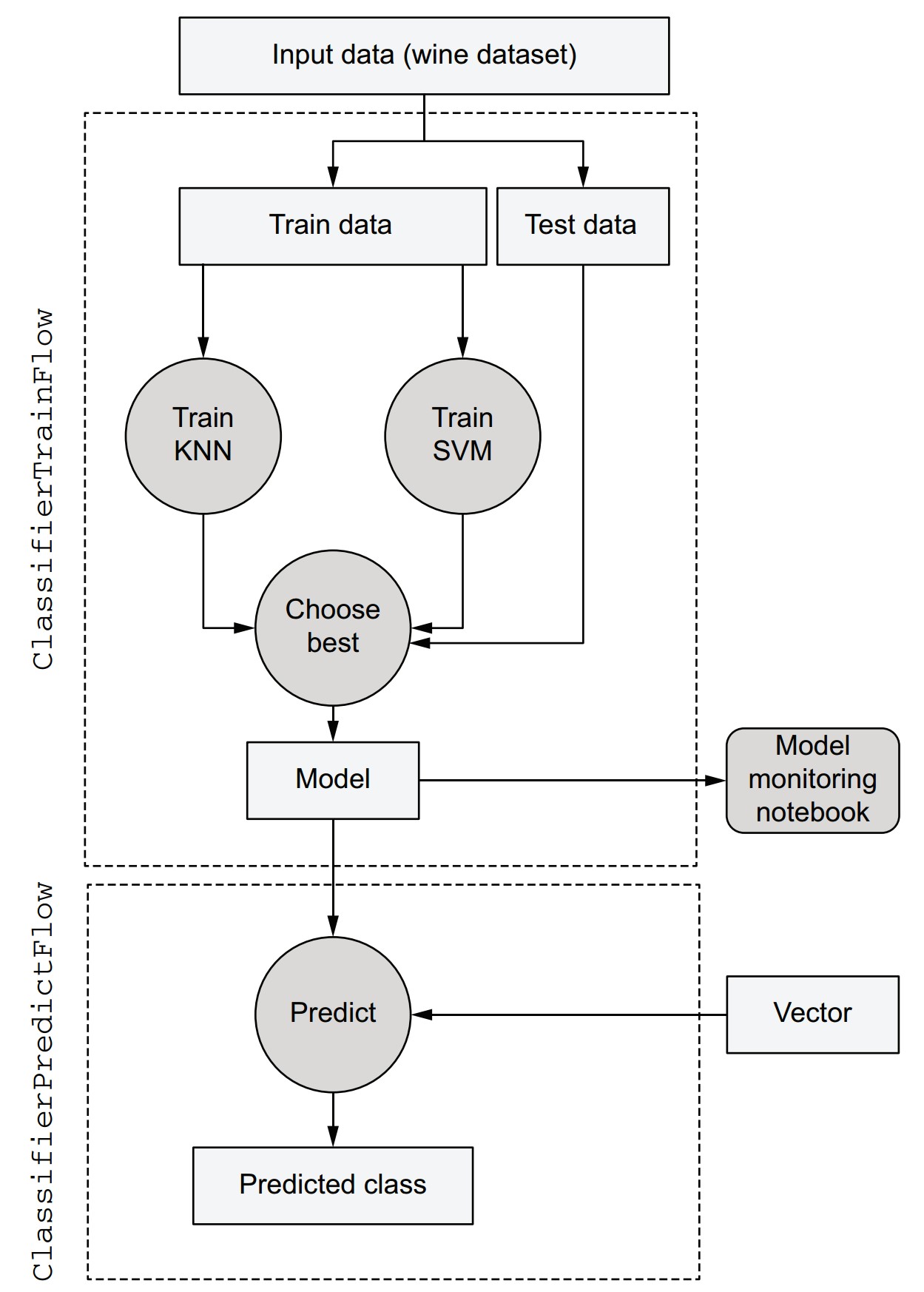
-
Input step:
- Loads and splits the dataset.
- Stores training/testing data as artifacts.
-
Parallel model training:
- Trains KNN and SVM in parallel steps.
-
Model selection:
- Evaluates both models using test data.
- Selects and stores the best model.
-
Artifact inspection:
- All intermediate and final data stored as artifacts.
- Accessible in notebooks via the Client API.
-
Prediction flow:
- Loads the latest model and classifies new data using the same interface.
Real-world scalability preview
This architecture scales to production scenarios. For example:
- Replace
--vectorwith a CSV file (IncludeFile). - Handle larger data inputs, multiple models, and long training times (e.g., with
foreachand parallel compute). - Use versioning, automation, and scheduling (covered in future chapters).
Questions answered in future
- How to handle large-scale datasets (TBs).
- How to scale up model training (e.g., 2,000 models).
- How to organize long flows across multiple files.
- How to schedule flows for production.
- How to isolate runs and use different dependencies per run.